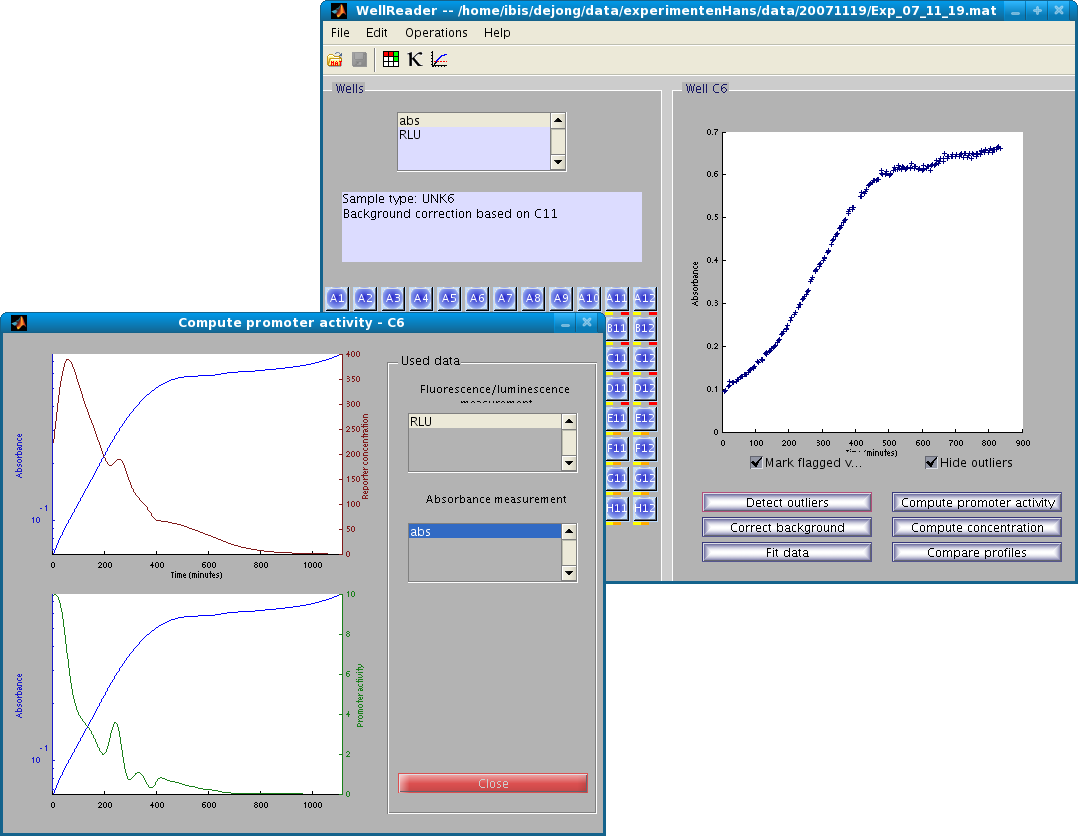
WellReader is a MATLAB application developed in the Ibis group (formerly the Helix group) at INRIA Grenoble - Rhône-Alpes. The software implements computational methods developed for the analysis of the dynamics of gene expression in microorganisms, measured by means of fluorescent and luminescent reporter genes.
As input, WellReader reads the primary data file produced by a 96-well microplate reader, containing the time-series measurements of the absorbance (optical density) levels as well as the fluorescence and luminescence levels in each well (if available).
The main view mimics the microplate, and various modules exist to analyze the data, in particular to detect outliers, correct data by background substraction, smooth the data, compute promoter activities and concentrations, compare expression profiles, etc.
At each step of the analysis, the results of the actions on the data are stored and can be saved and reloaded for further analysis.
In order to understand the function of the WellReader modules, and the way they combine in the analysis of the reporter gene data, have a look at the user guide, and then switch to the modules you are interested in.
Learn how to install and use WellReader with the user guide. We also feature a tutorial to come to grips with the tool.
See the primary data file and the WellReader file
See Contributors
See TechDoc.html